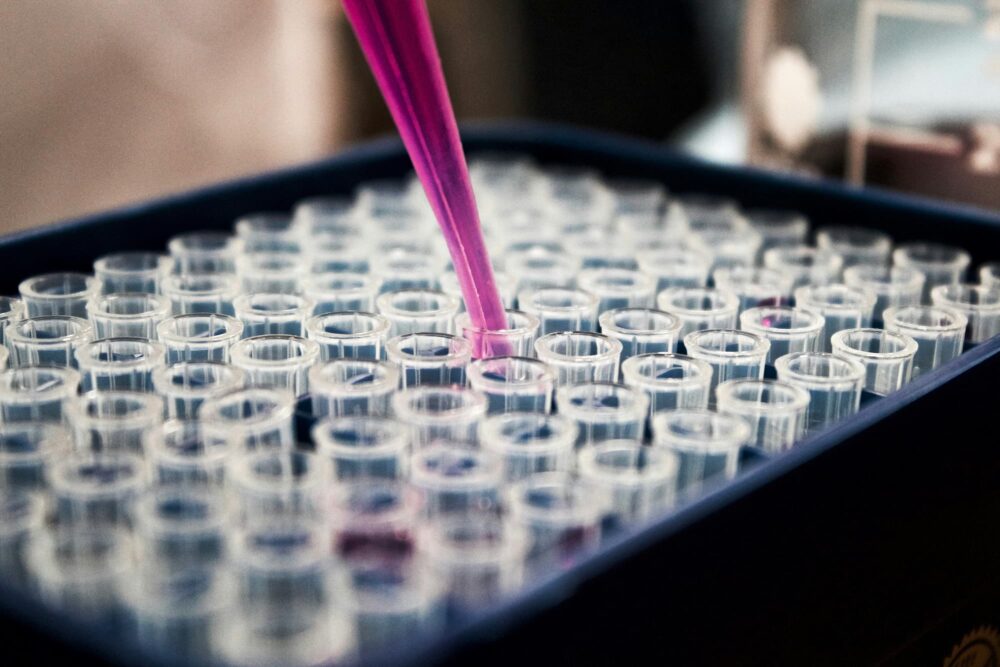
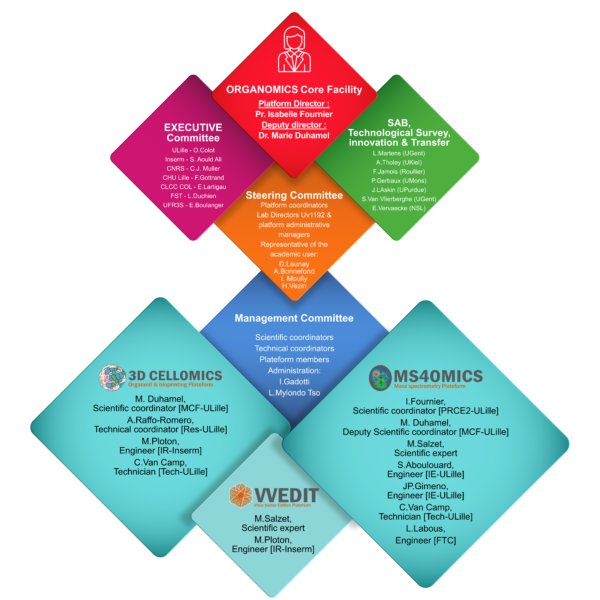
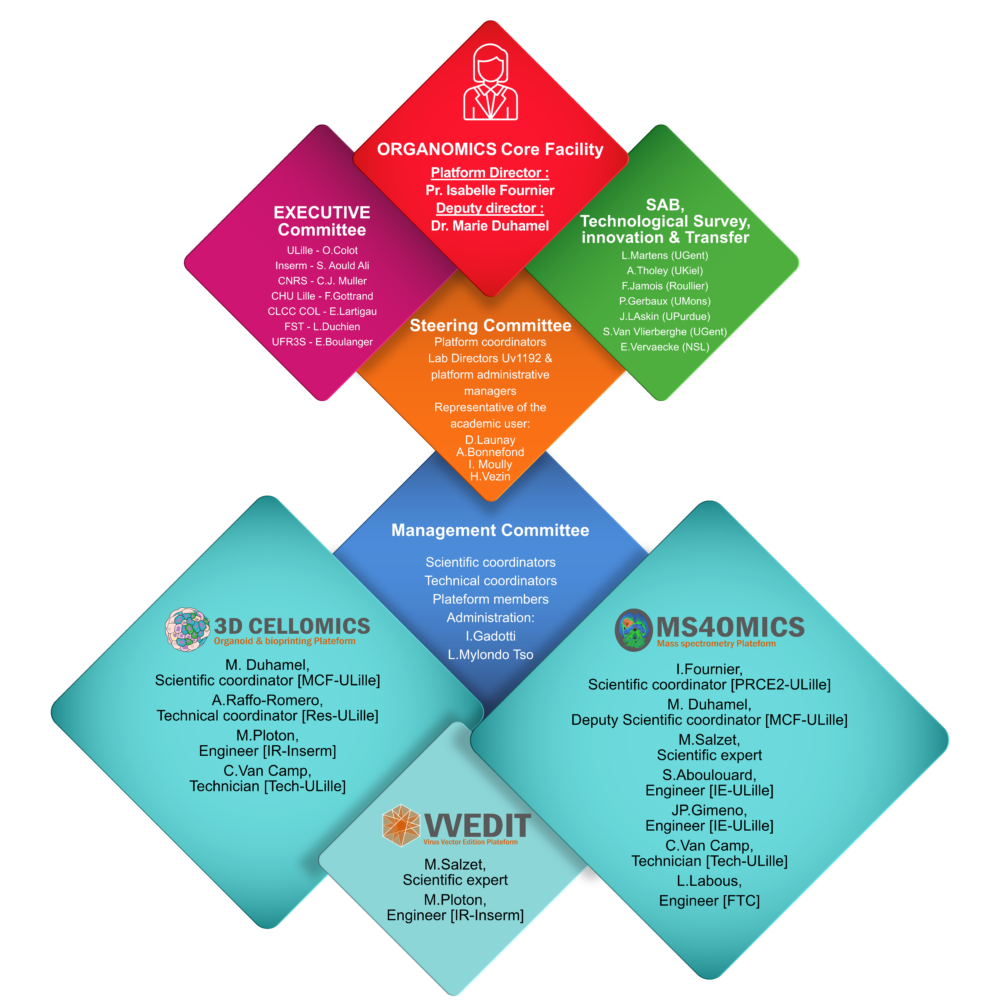
The ORGANOMICS platform of the PRISM Inserm U1192 laboratory offers expertise in proteomics, mass spectrometry imaging and the creation of 3D cellular models. In proteomics, it specializes in non-targeted approaches (bottom-up, shotgun and top-down) and interactomics (cross-linking MS and BioID). Particular expertise has been developed in spatial proteomics and FFPE tissue analysis, proteogenomics and exosome analysis. With regard to cellular models, the platform specializes in the creation of 3D models such as spheroids, organoids and bioprints, to mimic physiopathological contexts as closely as possible, which can be used to test new drugs, study intercellular communications and therapeutic resistance.
The platform is not open access, if you wish to use it please contact our teams to discuss the project and set up access to machines and techniques.